ASPL Operations Guide v 1.00
© 2025 Bassem W. Jamaleddine
8. 12f%U\ sequence alignment on the union of the mediated elements difference
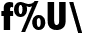
f%U\ a1 a2 a3 where a1 a2 a3 are set variables
f%U\`mtm~ a1 a2 a3
f%U\/mtm~ a1 a2 a3
seqop seqoid 3 notcommu ins:H_H_H text:alignment
The operator f%U\ is followed by three set variables. Use the operator f%U\ to align the union of two datasets where the first dataset is the result of the set difference of the first set variable with the third set variable, and the second dataset is the difference of the second set variable with the third set variable. It is assumed that each set variable has one group label, otherwise the first group (in the set variable) is explicitly implied. You can change the sequence alignment algorithm selectively. The sequence alignment can be selected: the command sequencing lcs selects the algorithm for the longest common sequence, and the command sequencing ssa selects the algorithm for the sequence similarity analysis.
f%U\ EXAMPLE1In this example, we will compare the JMX in Jar files in two directories by displaying their sequence alignment. We will start ASPL by loading the JMXJAR workspace, then we will issue the command f%U\ to display the alignment of the elements (which are the files) between the two set variables jmx1 and jmx2. The set variable jmx3 is mediated (using the difference set operation) with jmx1 and jmx2 prior to the alignment operation.
f%U\ OPERATION1# aspl JMXJAR
(start ASPL loading the sample workspace JMXJAR)
① aspl>
ks mtime chksum ppdd ffl
(set the ks vector to mtime chksum ppdd ffl)
② aspl>
f%U\ jmx1 jmx2 jmx3
(show the alignment of the elements union of "jmx1 \ jmx3" and "jmx2 \ jmx3")
③ aspl>
f%U\`c= jmx1 jmx2 jmx3
(show the alignment of the elements union of "jmx1 \ jmx3" and "jmx2 \ jmx3" such that checksums are equal)
④ aspl>
f%U\`c~ jmx1 jmx2 jmx3
(show the alignment of the elements union of "jmx1 \ jmx3" and "jmx2 \ jmx3" such that checksums are unequal)
NOTE: Here we mediate jmx3 and jmx1 with jmx2
⑤ aspl>
f%U\ jmx3 jmx1 jmx2
(show the alignment of the elements union of "jmx3 \ jmx2" and "jmx1 \ jmx2")
⑥ aspl>
f%U\`c= jmx3 jmx1 jmx2
(show the alignment of the elements union of "jmx3 \ jmx2" and "jmx1 \ jmx2" such that checksums are equal)
⑦ aspl>
f%U\`c~ jmx3 jmx1 jmx2
(show the alignment of the elements union of "jmx3 \ jmx2" and "jmx1 \ jmx2" such that checksums are unequal)
f%U& EXAMPLE2In this example, we will compare the JDBC Informix Jar files extracted in two directories by displaying their sequence alignment. We will start ASPL by loading the JDBCINFORMIX workspace, then we will issue the command f%U& to display the alignment of the elements (which are the files) between the two set variables ifxjdbc1 and ifxjdbc2. The set variable ifxjdbc3 is mediated (using the intersection operation) with ifxjdbc1 and ifxjdbc3 prior to the alignment operation.
f%U& OPERATION2# aspl JDBCINFORMIX
(start ASPL loading the sample workspace JDBCINFORMIX)
① aspl>
f%U\`mtm~ ifxjdbc1 ifxjdbc2 ifxjdbc3
(show the alignment of the elements union of "ifxjdbc1 & ifxjdbc3" and "ifxjdbc2 & ifxjdbc3" such that mtimes are unequal)
② aspl>
f%U\`c= ifxjdbc1 ifxjdbc2 ifxjdbc3
(show the alignment of the elements union of "ifxjdbc1 \ ifxjdbc3" and "ifxjdbc2 \ ifxjdbc3" such that checksums are equal)
③ aspl>
f%U\`c~ ifxjdbc1 ifxjdbc2 ifxjdbc3
(show the alignment of the elements union of "ifxjdbc1 & ifxjdbc3" and "ifxjdbc2 & ifxjdbc3" such that checksums are unequal)
Here we mediate ifxjdbc1 and ifxjdbc3 with the ifxjdbc2
④ aspl>
f%U\`c~ ifxjdbc1 ifxjdbc3 ifxjdbc2
(show the alignment of the elements union of "ifxjdbc1 & ifxjdbc2" and "ifxjdbc3 & ifxjdbc2" such that checksums are unequal)
● Tickable Predicates for f%U\
This f%U\ is tickable with predicates that are shown in the following table:
f%U\`_ sequence alignment on the union of the mediated elements difference f%U\`c= sequence alignment on the union of the mediated elements difference such that they have the same checksum f%U\`c~ sequence alignment on the union of the mediated elements difference such that they have different checksums f%U\`c~~ sequence alignment on the union of the mediated elements difference such that they have all over different checksums f%U\`h= sequence alignment on the union of the mediated elements difference such that they have the same entropy f%U\`h~ sequence alignment on the union of the mediated elements difference such that they have different entropies f%U\`h~~ sequence alignment on the union of the mediated elements difference such that they have all over different entropies f%U\`mtm= sequence alignment on the union of the mediated elements difference such that they have the same make time f%U\`mtm~ sequence alignment on the union of the mediated elements difference such that they have different make times f%U\`mtm~~ sequence alignment on the union of the mediated elements difference such that they have all over different make times f%U\`mtm< sequence alignment on the union of the mediated elements difference such that they have a prior make time f%U\`mtm<= sequence alignment on the union of the mediated elements difference such that they have prior or same make time f%U\`mtm> sequence alignment on the union of the mediated elements difference such that they have more recent make time f%U\`mtm>= sequence alignment on the union of the mediated elements difference such that they have more recent or same make time f%U\`ks= sequence alignment on the union of the mediated elements difference such that they have the same ksum f%U\`ks~ sequence alignment on the union of the mediated elements difference such that they have different ksums f%U\`ks~~ sequence alignment on the union of the mediated elements difference such that they have all over different ksums